Working with images#
To do image data analysis, we first need to be able to do a few essential operations:
opening images
displaying images
have a look at pixel statistics
See also
Opening images#
Most images with standard extensions (tif, png etc.) can be read using the skimage.io.imread function. In case your image doesn’t you should consult the documentation of the given file format.
To use imread you have three possibilities:
use the absolute path to an image file e.g.
imread('/Users/username/Desktop/blobs.tif')use a relative path to where you currently are (you cand find out using the
pwdcommand in a cell) i.e.imread('../../data/blobs.tif')use a url that points to an image file, for example from the GitHub repository
imread('https://github.com/haesleinhuepf/BioImageAnalysisNotebooks/raw/main/data/blobs.tif')
Here we use a relative path. Respective to the current notebook, the data are two folder levels higher (../../) in a folder called data:
from skimage.io import imread
image = imread("../../data/blobs.tif")
As shown earlier, images are just matrices of intensities. However, showing them as such is not convenient.
image
array([[ 40, 32, 24, ..., 216, 200, 200],
[ 56, 40, 24, ..., 232, 216, 216],
[ 64, 48, 24, ..., 240, 232, 232],
...,
[ 72, 80, 80, ..., 48, 48, 48],
[ 80, 80, 80, ..., 48, 48, 48],
[ 96, 88, 80, ..., 48, 48, 48]], dtype=uint8)
Displaying images#
There are many ways to display simple 2D images. In many notebooks and examples online, you will find examples using Matplotlib’s imshow function on an array:
from matplotlib import pyplot as plt
plt.imshow(image);
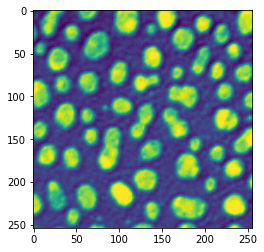

Unfortunately, the imshow function is not an optimal choice to display microscopy images: it doesn’t handle well multi-channel data, it is difficult to handle intensity ranges etc. Throughout this course we therefore favor the use of the microshow function from the microfilm package or the imshow function from the clesperanto package. We’ll learn about the second solution in coming chapters. Here we just use microshow:
from microfilm.microplot import microshow
microshow(image);
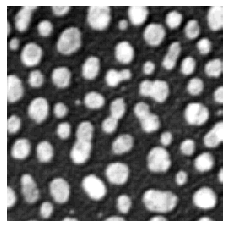

Lookup tables (a.k.a. color maps)#
We can also change the look-up table, a.k.a. “color map” for the visualization.
microshow(image, cmaps="hot");
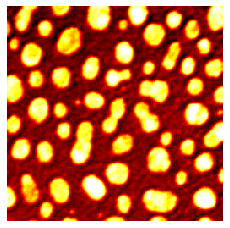

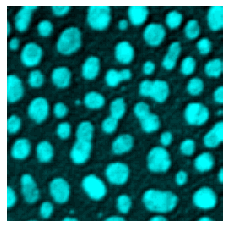
microshow(image, cmaps="pure_cyan");
Exercise#
Open the banana020.tif data set, visualize it in a yellowish lookup table.
